Page 187 - 《广西植物》2026年第1期
P. 187
1 期 王宇等: 濒危植物珙桐根际微生物和根内生菌群落特点及功能分析 1 8 3
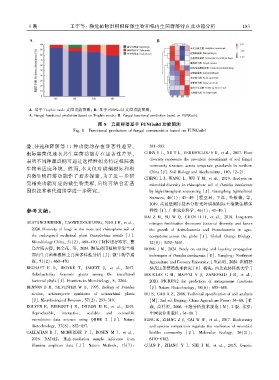
A. 基于 Trophic mode 真菌功能预测ꎻ B. 基于 FUNGuild 真菌功能预测ꎮ
A. Fungal functional prediction based on Trophic modeꎻ B. Fungal functional prediction based on FUNGuild.
图 8 真菌群落基于 FUNGuild 功能预测
Fig. 8 Functional prediction of fungal communities based on FUNGuild
叠、分选和降解等 11 种功能均存在显著性差异ꎬ 581-583.
CHEN Y Lꎬ XU T Lꎬ VERESOGLOU S Dꎬ et al.ꎬ 2017. Plant
根际真菌仅地衣共生真菌功能存在显著性差异ꎬ
表明不同种源珙桐可通过选择性招募特定根际微 diversity represents the prevalent determinant of soil fungal
community structure across temperate grasslands in northern
生物来适应环境ꎮ 然而ꎬ本文仅对珙桐根际和根
China [J]. Soil Biology and Biochemistryꎬ 110: 12-21.
内微生物群落功能作了初步探索ꎬ为了进一步研
CHENG L Jꎬ WANG Lꎬ WU Y Mꎬ et al.ꎬ 2019. Analysis on
究相关功能对应的微生物类群ꎬ后续可结合宏基 microbial diversity in rhizosphere soil of Davidia involucrate
因组技术和代谢组学进一步研究ꎮ by high ̄throughput sequencing [J]. Guangdong Agricultural
Sciencesꎬ 46(1): 43-49. [程立君ꎬ 王磊ꎬ 吴银梅ꎬ 等ꎬ
2019. 高通量测序技术分析光叶珙桐根际土壤微生物多
参考文献: 样性 [J]. 广东农业科学ꎬ 46(1): 43-49.]
DAI Z Mꎬ SU W Qꎬ CHEN H Hꎬ et al.ꎬ 2018. Long ̄term
ALATANCUMBUERꎬ CAOWUJISIGULENGꎬ BAO J Hꎬ et al.ꎬ nitrogen fertilization decreases bacterial diversity and favors
2024. Diversity of fungi in the roots and rhizosphere soil of the growth of Actinobacteria and Proteobacteria in agro ̄
the endangered medicinal plant Dactylorhiza viridis [ J]. ecosystems across the globe [ J]. Global Change Biologyꎬ
Microbiology Chinaꎬ 51(2): 460-470. [阿拉坦存布尔ꎬ 曹 24(8): 3452-3461.
乌吉斯古楞ꎬ 包金花ꎬ 等ꎬ 2024. 濒危药用植物掌裂兰根 DONG J Mꎬ 2024. Study on cutting and layering propagation
部内生真菌和根际土真菌多样性分析 [J]. 微生物学通 techniques of Davidia involucrata [D]. Yangling: Northwest
报ꎬ 51(2): 460-470. Agriculture and Forestry University. [董嘉明ꎬ 2024. 珙桐扦
BECRAFT E Dꎬ WOYKE Tꎬ JARETT Jꎬ et al.ꎬ 2017. 插及压条繁殖技术研究 [D]. 杨凌: 西北农林科技大学.]
Rokubacteria: Genomic giants among the uncultured DOUGLAS G Mꎬ MAFFEI V Jꎬ ZANEVELD J Rꎬ et al.ꎬ
bacterial phyla [J]. Frontiers in Microbiologyꎬ 8: 2264. 2020. PICRUSt2 for prediction of metagenome functions
BENSON D Rꎬ SILVESTER W Bꎬ 1993. Biology of Frankia [J]. Nature Biotechnologyꎬ 38(6): 685-688.
strainsꎬ actinomycete symbionts of actinorhizal plants DU Sꎬ GAO X Zꎬ 2006. Technical specification of soil analysis
[J]. Microbiological Reviewsꎬ 57(2): 293-319. [M]. 2nd ed. Beijing: China Agriculture Press: 34-68. [杜
BOLYEN Eꎬ RIDEOUT J Rꎬ DILLON M Rꎬ et al.ꎬ 2019. 森ꎬ 高祥照ꎬ 2006. 土壤分析技术规范 [M]. 2 版. 北京:
Reproducibleꎬ interactiveꎬ scalable and extensible 中国农业出版社: 34-68.]
microbiome data science using QIIME 2 [ J ]. Nature FENG Kꎬ ZHANG Z Jꎬ CAI W Wꎬ et al.ꎬ 2017. Biodiversity
Biotechnologyꎬ 37(8): 852-857. and species competition regulate the resilience of microbial
CALLAHAN B Jꎬ MCMURDIE P Jꎬ ROSEN M Jꎬ et al.ꎬ biofilm community [ J ]. Molecular Ecologyꎬ 26(21):
2016. DADA2: High ̄resolution sample inference from 6170-6182.
illumina amplicon data [ J ]. Nature Methodsꎬ 13(7): GUAN Pꎬ ZHANG Y Jꎬ SHI J Mꎬ et al.ꎬ 2015. Genetic